
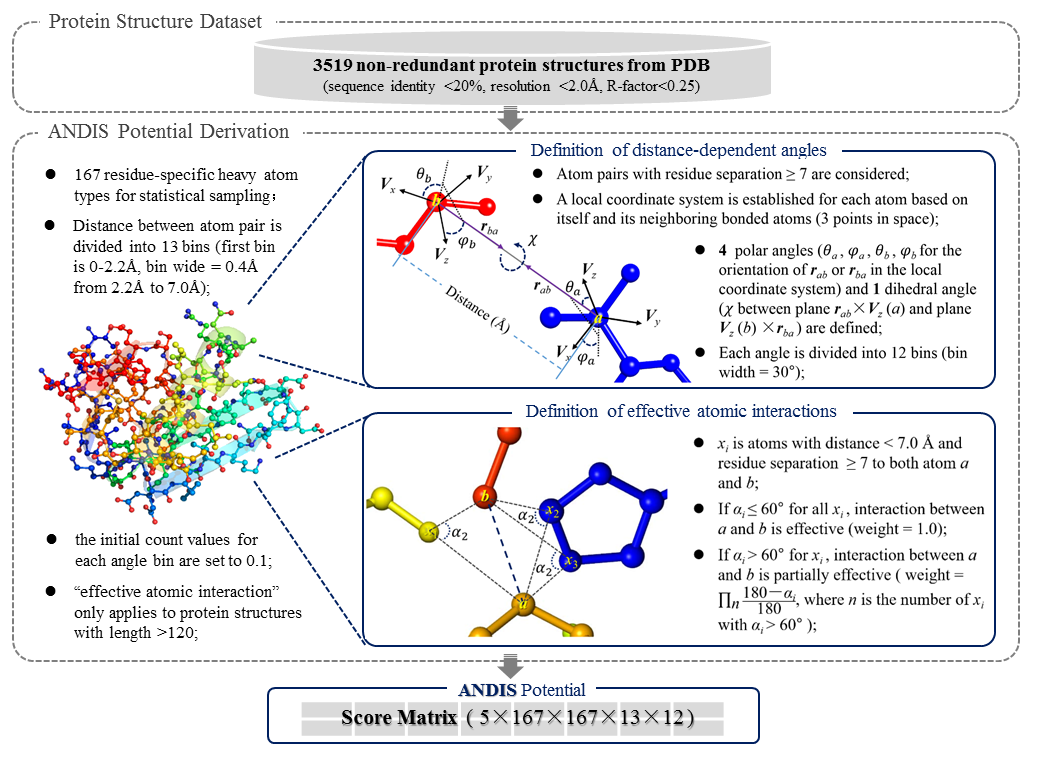
ANDIS is an atomic angle- and distance-dependent statistical potential devoted for protein structure quality assessment. Five angles are calculated for each atom-pair with distance < dcut (dcut is adjustable from 7.0 to 15.0 angstrom) and residue separation ≥ 7 in protein structure. the observed frequencies of atom-pair distances and corresponding angles are converted to a combined statistical potential based on the inverse boltzmann law. andis naturally integrates the atomic orientation-dependent and distance-dependent interactions. we benchmarked andis with a comprehensive list of publicly available statistical potentials. andis demonstrate a significantly better ability of native recognition than the other tested potentials.
ANDIS Resource
Download ANDIS standalone program: ANDIS.zip (95MB)
Download non-redundant PDB dataset (3519 experimental structures with Identity cutoff =20%, Resolution cutoff =2.0 angstroms, R-factor cutoff =0.25): PDBdataset.zip (115MB)
Download 175 CASP10-13 decoy sets: CASP10-13.zip (354MB)
Reference:

